How to Use map2() and pmap() in R
Introduction
While map() iterates over a single list, map2() iterates over two lists in parallel, and pmap() iterates over any number of lists. These functions are essential when your operation needs inputs from multiple sources.
When to use: - map2() - when you have exactly two parallel inputs - pmap() - when you have three or more parallel inputs, or a data frame of parameters
Getting Started
library(tidyverse)map2(): Iterate Over Two Lists
Basic syntax
map2() takes two lists and applies a function using corresponding elements:
xs <- list(1, 2, 3)
ys <- list(10, 20, 30)
# Add corresponding elements
map2(xs, ys, \(x, y) x + y)Practical example: weighted means
values <- list(
a = c(1, 2, 3),
b = c(10, 20, 30),
c = c(100, 200, 300)
)
weights <- list(
a = c(0.5, 0.3, 0.2),
b = c(0.1, 0.2, 0.7),
c = c(0.33, 0.33, 0.34)
)
# Calculate weighted mean for each pair
map2_dbl(values, weights, weighted.mean)String operations with two inputs
first_names <- c("John", "Jane", "Bob")
last_names <- c("Doe", "Smith", "Johnson")
# Combine names
map2_chr(first_names, last_names, \(f, l) paste(f, l))
# Or using paste directly
map2_chr(first_names, last_names, paste)Creating file paths
folders <- c("data", "data", "output")
files <- c("input.csv", "config.json", "results.csv")
map2_chr(folders, files, file.path)map2() Variants
Just like map(), map2() has typed variants:
xs <- 1:3
ys <- 4:6
map2_dbl(xs, ys, `+`) # numeric: 5, 7, 9
map2_int(xs, ys, `+`) # integer: 5L, 7L, 9L
map2_chr(xs, ys, paste) # character: "1 4", "2 5", "3 6"
map2_lgl(xs, ys, `<`) # logical: TRUE, TRUE, TRUEpmap(): Iterate Over Multiple Lists
Basic syntax
pmap() takes a list of lists (or a data frame) and applies a function:
params <- list(
n = c(10, 20, 30),
mean = c(0, 5, 10),
sd = c(1, 2, 3)
)
# Generate random samples with different parameters
set.seed(42)
pmap(params, rnorm)Using a data frame as input
Data frames are naturally suited for pmap():
# Create parameter grid
param_grid <- expand_grid(
n = c(100, 1000),
mean = c(0, 10),
sd = c(1, 5)
)
param_grid
# Generate samples for each row
set.seed(42)
samples <- pmap(param_grid, rnorm)
map_dbl(samples, mean) # Check the meansNamed arguments matter
pmap() matches list names to function arguments:
# Names must match function argument names
params <- list(
x = c(1, 2, 3),
y = c(4, 5, 6),
z = c(7, 8, 9)
)
# Custom function
my_sum <- function(x, y, z) x + y + z
pmap_dbl(params, my_sum)Practical Example: Model Comparison
Run multiple models with different specifications
# Define model specifications
model_specs <- tibble(
formula = c(
"mpg ~ wt",
"mpg ~ wt + hp",
"mpg ~ wt + hp + am"
),
data = list(mtcars, mtcars, mtcars)
)
# Fit all models
models <- pmap(model_specs, \(formula, data) {
lm(as.formula(formula), data = data)
})
# Extract R-squared values
map_dbl(models, \(m) summary(m)$r.squared)Create multiple plots
# Parameters for plots
plot_params <- tibble(
x_var = c("wt", "hp", "disp"),
y_var = rep("mpg", 3),
title = c("Weight vs MPG", "Horsepower vs MPG", "Displacement vs MPG")
)
# Generate plots (returns list of ggplots)
plots <- pmap(plot_params, \(x_var, y_var, title) {
ggplot(mtcars, aes(x = .data[[x_var]], y = .data[[y_var]])) +
geom_point() +
geom_smooth(method = "lm") +
labs(title = title) +
theme_minimal()
})
# Display first plot
plots[[1]]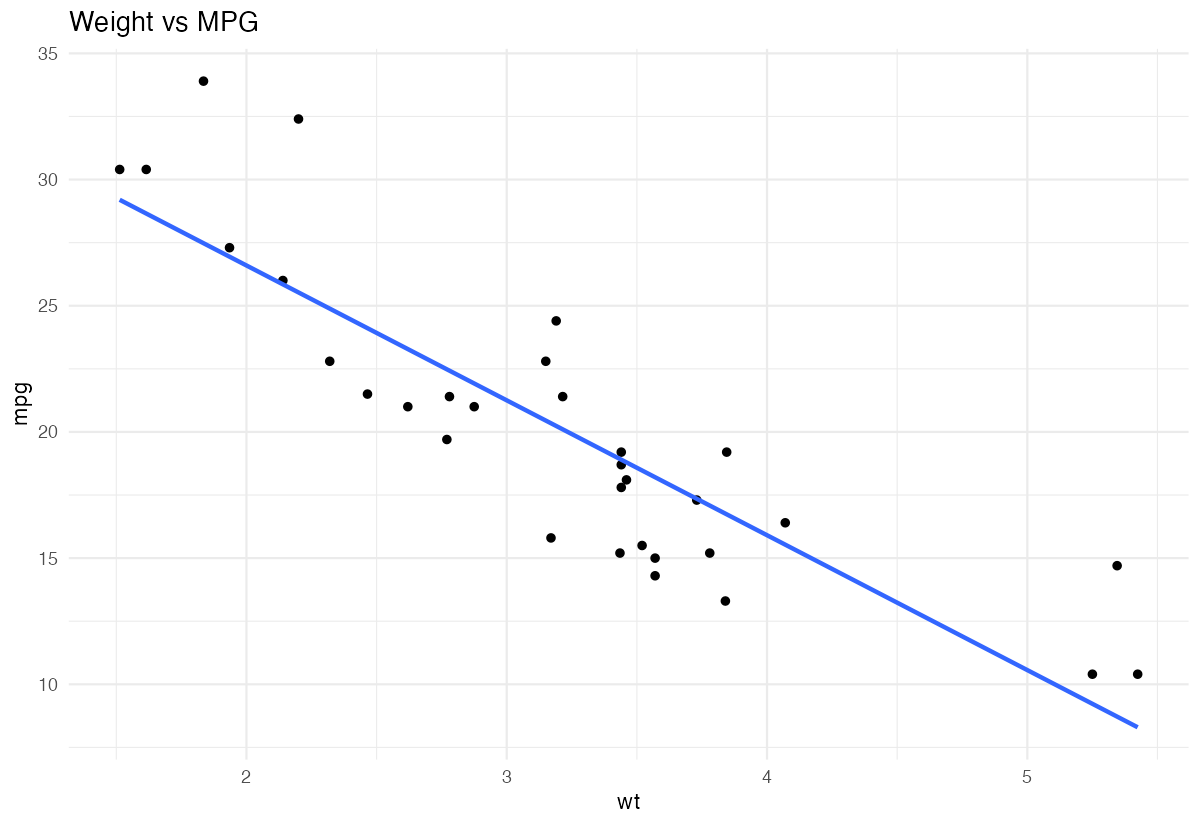
imap(): Iterate with Index or Name
When you need both the element AND its index/name, use imap():
x <- c(a = 10, b = 20, c = 30)
# imap passes element and name/index
imap_chr(x, \(value, name) paste(name, "=", value))
# "a = 10", "b = 20", "c = 30"With unnamed vectors, you get the index
letters[1:3] |>
imap_chr(\(letter, idx) paste0(idx, ": ", letter))
# "1: a", "2: b", "3: c"Practical use: progress messages
files <- c("data1.csv", "data2.csv", "data3.csv")
iwalk(files, \(file, i) {
message(sprintf("[%d/%d] Processing %s", i, length(files), file))
# read_csv(file) ...
})Add row numbers to nested data
penguins |>
drop_na() |>
split(~species) |>
imap(\(df, species_name) {
df |> mutate(species_label = species_name)
}) |>
list_rbind()Performance: pmap vs Nested Loops
pmap() is cleaner and often faster than nested loops:
# Parameter grid
params <- expand_grid(a = 1:10, b = 1:10, c = 1:10)
# Instead of nested loops:
# for (a in 1:10) {
# for (b in 1:10) {
# for (c in 1:10) { ... }
# }
# }
# Use pmap:
results <- pmap_dbl(params, \(a, b, c) a * b + c)This is more readable and avoids growing vectors in loops.
walk2() and pwalk(): For Side Effects
When you want side effects (like saving files) instead of return values:
# Save multiple datasets
datasets <- list(
iris = iris,
mtcars = mtcars
)
filenames <- c("iris.csv", "mtcars.csv")
# This would write files (commented out)
# walk2(datasets, filenames, \(data, file) {
# write_csv(data, file)
# })Common Mistakes
1. Lists of different lengths
# Shorter list is NOT recycled (unlike base R)
xs <- 1:3
ys <- 1:2
# This errors:
# map2(xs, ys, `+`)
# Make lengths match explicitly
map2(xs, c(ys, NA), `+`)2. Forgetting to name pmap arguments
# This might not work as expected if names don't match
params <- list(c(10, 20), c(0, 5), c(1, 2)) # unnamed
# Better: use names that match function arguments
params <- list(n = c(10, 20), mean = c(0, 5), sd = c(1, 2))
pmap(params, rnorm)3. Using map2 when you need pmap
# For more than 2 inputs, use pmap
# Wrong approach: nested map2
# Right approach: pmap with all inputsSummary
| Function | Inputs | Returns | Use Case |
|---|---|---|---|
map2() |
2 lists | list | Combine two parallel sequences |
map2_dbl() |
2 lists | numeric | Two inputs, numeric output |
pmap() |
n lists | list | Multiple parameters, complex operations |
pmap_dfr() |
n lists | data frame | Parameter grid → combined results |
walk2() |
2 lists | invisible | Two inputs, side effects only |
pwalk() |
n lists | invisible | Multiple inputs, side effects |
- Use
map2()for exactly two parallel inputs - Use
pmap()for three or more inputs, or when working with data frames - Name your list elements to match function argument names
- Use
walk2()/pwalk()for side effects like saving files